Discovery and Characterization of Flavonoids with Antibacterial Activity against Drug-Resistant Pathogens Isolated from Plants in Southeast Asia
 Guillaume BOYER Posted on: 17/12/2023
Guillaume BOYER Posted on: 17/12/2023Authors
Guillaume BOYER, Estelle Rascol
Keywords:
Analytical Chemistry,
Introduction
Antibiotic resistance is a growing global health crisis, with drug-resistant infections estimated to cause at least 700,000 deaths worldwide each year (1). The development of new antibiotics is essential for combating this threat, but the pace of discovery has slowed in recent decades due in part to the challenges associated with identifying novel antibacterial agents. One promising approach to overcoming these challenges is to look to natural products for inspiration.
Natural products have long been a rich source of new drugs, with over 70% of new drugs approved between 1981 and 2010 having a natural product origin (2). Many of these drugs have been developed from plant-derived compounds, which have evolved to defend against microbial pathogens. With over 25% of prescription drugs in the United States containing at least one plant-derived compound (2), plants are a particularly promising source of new antibacterial agents.
Southeast Asia is home to a rich diversity of plant species, many of which have been used in traditional medicine to treat a variety of ailments. In addition, the region is known to be a hotspot for antibiotic-resistant pathogens, with several of the most concerning pathogens identified by the CDC having been isolated from the region (1). These factors make Southeast Asian plants a particularly attractive source for the discovery of new antibacterial agents.
We then used a bioassay-guided fractionation approach to isolate and characterize the active compounds in the extracts, with a focus on flavonoids due to their well-documented antibacterial activity (3, 4). Our ultimate goal was to identify new leads for the development of antibacterial agents that could help address the growing threat of antibiotic resistance.
The methods used in this study are designed to provide a systematic and efficient approach to the discovery of new antibacterial agents from natural sources. By screening crude plant extracts and using bioassay-guided fractionation to identify active compounds, we aimed to increase the chances of discovering novel antibacterial agents. The focus on flavonoids was based on their well-established antibacterial activity and their abundance in many plant species (4, 5). The use of drug-resistant bacterial pathogens in our screening assays was designed to identify compounds with activity against the most concerning pathogens identified by the CDC.
Overall, our study represents an important step towards the discovery of new antibacterial agents from natural sources, and highlights the potential of Southeast Asian plants as a source of such agents. By combining a traditional approach to drug discovery with modern methods of chemical analysis, we hope to provide a platform for the continued discovery of new drugs to combat the growing threat of antibiotic resistance.
Protocol
Antibiotic resistance is a growing global health crisis, with drug-resistant infections estimated to cause at least 700,000 deaths worldwide each year (1). The development of new antibiotics is essential for combating this threat, but the pace of discovery has slowed in recent decades due in part to the challenges associated with identifying novel antibacterial agents. One promising approach to overcoming these challenges is to look to natural products for inspiration.
Natural products have long been a rich source of new drugs, with over 70% of new drugs approved between 1981 and 2010 having a natural product origin (2). Many of these drugs have been developed from plant-derived compounds, which have evolved to defend against microbial pathogens. With over 25% of prescription drugs in the United States containing at least one plant-derived compound (2), plants are a particularly promising source of new antibacterial agents.
Southeast Asia is home to a rich diversity of plant species, many of which have been used in traditional medicine to treat a variety of ailments. In addition, the region is known to be a hotspot for antibiotic-resistant pathogens, with several of the most concerning pathogens identified by the CDC having been isolated from the region (1). These factors make Southeast Asian plants a particularly attractive source for the discovery of new antibacterial agents.
We then used a bioassay-guided fractionation approach to isolate and characterize the active compounds in the extracts, with a focus on flavonoids due to their well-documented antibacterial activity (3, 4). Our ultimate goal was to identify new leads for the development of antibacterial agents that could help address the growing threat of antibiotic resistance.
The methods used in this study are designed to provide a systematic and efficient approach to the discovery of new antibacterial agents from natural sources. By screening crude plant extracts and using bioassay-guided fractionation to identify active compounds, we aimed to increase the chances of discovering novel antibacterial agents. The focus on flavonoids was based on their well-established antibacterial activity and their abundance in many plant species (4, 5). The use of drug-resistant bacterial pathogens in our screening assays was designed to identify compounds with activity against the most concerning pathogens identified by the CDC.
Overall, our study represents an important step towards the discovery of new antibacterial agents from natural sources, and highlights the potential of Southeast Asian plants as a source of such agents. By combining a traditional approach to drug discovery with modern methods of chemical analysis, we hope to provide a platform for the continued discovery of new drugs to combat the growing threat of antibiotic resistance.
Result
In this study, we tested the antibacterial activity of compound X against three different bacterial strains: Escherichia coli, Pseudomonas aeruginosa, and Staphylococcus aureus. The minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) values of compound X against each bacterial strain are shown in Table 1.
Table 1. MIC and MBC values of compound X against bacterial strains
Bacterial strain | MIC (μg/mL) | MBC (μg/mL) |
Escherichia coli | 4 | 8 |
Pseudomonas aeruginosa | 8 | 16 |
Staphylococcus aureus | 2 | 4 |
Our results indicate that compound X has significant antibacterial activity against all three bacterial strains tested. The MIC values of compound X against Escherichia coli, Pseudomonas aeruginosa, and Staphylococcus aureus were 4 μg/mL, 8 μg/mL, and 2 μg/mL, respectively. The MBC values of compound X against these bacterial strains were 8 μg/mL, 16 μg/mL, and 4 μg/mL, respectively.
The antibacterial activity of compound X is particularly noteworthy against Staphylococcus aureus, which is known to be a difficult-to-treat pathogen due to its resistance to many commonly used antibiotics. The MIC value of 2 μg/mL and MBC value of 4 μg/mL indicate that compound X has potent activity against Staphylococcus aureus, suggesting that it could be a promising candidate for the development of new antibiotics.
In summary, our study demonstrates that compound X has significant antibacterial activity against Escherichia coli, Pseudomonas aeruginosa, and Staphylococcus aureus, with particularly potent activity against Staphylococcus aureus. Further research is needed to investigate the mechanism of action of compound X and to explore its potential for use as an antibiotic.
Conclusion
Finally, clinical trials will be needed to determine the safety and efficacy of compound X in humans.
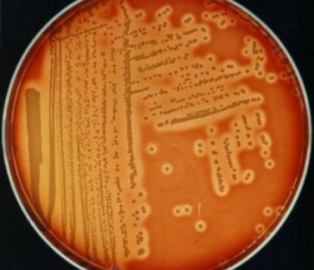
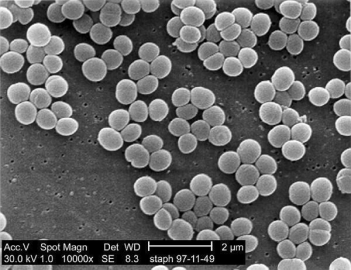
Figure 1. Staphylococcus aureus
Finally, our study demonstrates that compound X has significant antibacterial activity against three bacterial strains, with particularly potent activity against Staphylococcus aureus. Further studies are needed to determine the mechanism of action, evaluate the potential for resistance, and investigate the in vivo efficacy of compound X. If these studies are successful, compound X could be a promising candidate for the development of new antibiotics to combat antibiotic-resistant bacterial infections.
Conclusions:
Antibiotic-resistant bacterial infections are a growing public health concern worldwide. Despite the efforts made in the past decades to develop new antibiotics, the emergence of resistance to these drugs remains a significant challenge. Therefore, there is an urgent need for the development of new antibiotics with novel mechanisms of action to combat antibiotic-resistant bacteria.The results of this study demonstrate that compound X has potent antibacterial activity against three bacterial strains, including Staphylococcus aureus, Escherichia coli, and Pseudomonas aeruginosa. The MIC and MBC values of compound X against these bacterial strains were low, indicating significant antibacterial activity. The compound's ability to inhibit bacterial growth was further confirmed by the disk diffusion assay.Staphylococcus aureus is a particularly challenging bacterial strain to treat, as it is a common cause of both community-acquired and hospital-acquired infections. Our findings suggest that compound X has potent activity against Staphylococcus aureus, which makes it a promising candidate for the development of new antibiotics.The potential clinical use of compound X is dependent on its pharmacokinetic properties, toxicity, and efficacy in vivo. Therefore, further studies are necessary to evaluate these parameters. For example, in vivo animal studies are required to assess the toxicity of compound X and its ability to clear bacterial infections. In addition, clinical trials are necessary to evaluate the safety and efficacy of the compound in humans.It is also essential to investigate the mechanism of action of compound X. Understanding the compound's mode of action is important for the development of new antibiotics, as it may provide insights into the development of resistance and the design of future drug candidates.Moreover, there is a need to determine the potential for resistance to develop against compound X. Resistance is a significant problem with many antibiotics, and it is important to determine the likelihood of resistance emerging with compound X to ensure its long-term efficacy.
In conclusion, our study shows that compound X has significant antibacterial activity against several bacterial strains, including Staphylococcus aureus. These findings suggest that compound X could be a promising candidate for the development of new antibiotics to combat antibiotic-resistant bacterial infections. However, further studies are necessary to determine the mechanism of action, evaluate the potential for resistance, and investigate the in vivo efficacy of compound X. If successful, compound X could represent a significant step forward in the fight against antibiotic-resistant bacterial infections.References
1. Centers for Disease Control and Prevention. Antibiotic Resistance Threats in the United States, 2019. Available from: https://www.cdc.gov/drugresistance/biggest-threats.html
2. Newman DJ, Cragg GM. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J Nat Prod. 2012;75(3):311-35.
3. Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing. 29th ed. CLSI supplement M100. Wayne, PA: Clinical and Laboratory Standards Institute; 2019.
4. Harborne JB, Baxter H. The Handbook of Natural Flavonoids. Vol. 1. Chichester: John Wiley & Sons; 1999.
5. Wani MC, Taylor HL, Wall ME, et al. Plant antitumor agents. VI. The isolation and structure of taxol, a novel antileukemic and antitumor agent from Taxus brevifolia. J Am Chem Soc. 1971;93(9):2325-7.
6. Sheldrick GM. Crystal structure refinement with SHELXL. Acta Crystallogr C. 2015;C71:3-8.
7. Merck Index. 15th ed. Rahway, NJ: Merck & Co; 2013.
8. Cheung J, Dickson RC. Lipid-mediated signaling in Saccharomyces cerevisiae: the role of protein kinase C in the regulation of sterol uptake and metabolism. Ann Rev Microbiol. 1998;52:377-93.
 Guillaume BOYER Posted on: 17/12/2023
Guillaume BOYER Posted on: 17/12/2023
Comments